|
| | PSSMOptEPositionData () |
| |
| virtual | ~PSSMOptEPositionData () |
| |
| void | set_pssm_probabilities (utility::vector1< Real > const &pssm_probs) |
| |
| virtual Real | get_score (Multivec const &component_weights, Multivec const &vars, Multivec &dE_dvars, Size const num_energy_dofs, int const num_ref_dofs, int const num_total_dofs, EnergyMap const &fixed_terms, ScoreTypes const &score_list, ScoreTypes const &fixed_score_list) const |
| |
| virtual void | print_score (std::ostream &ostr, Multivec const &component_weights, Multivec const &vars, Multivec &dE_dvars, Size const num_energy_dofs, int const num_ref_dofs, int const num_total_dofs, EnergyMap const &fixed_terms, ScoreTypes const &score_list, ScoreTypes const &fixed_score_list) const |
| |
| virtual OptEPositionDataType | type () const |
| |
| virtual void | write_to_file (std::ofstream &outfile) const |
| |
| virtual void | read_from_file (std::ifstream &infile) |
| |
| virtual void | write_to_binary_file (std::ofstream &outfile) const |
| |
| virtual void | read_from_binary_file (std::ifstream &infile) |
| |
| virtual Size | memory_use () const |
| |
| | PNatAAOptEPositionData () |
| |
| virtual | ~PNatAAOptEPositionData () |
| |
| virtual void | range (ScoreTypes const &free_score_list, ScoreTypes const &fixed_score_list, EnergyMap &lower_bound, EnergyMap &upper_bound) const |
| | Return the upper and lower bound on the unweighted components at this position if they are larger (or smaller) than the unweighted values already in the two input EnergyMaps. More...
|
| |
| virtual Size | size () const |
| |
| void | set_position (Size pos_in) |
| |
| Size | position () const |
| |
| void | set_native_aa (AA nat_in) |
| |
| AA | native_aa () const |
| |
| void | set_neighbor_count (Size nb_in) |
| |
| Size | neighbor_count () const |
| |
| void | add_rotamer_line_data (PNatAAOptERotamerDataOP rot_in) |
| |
| PNatAAOptERotamerDataOPs & | data () |
| |
| PNatAAOptERotamerDataOPs const & | data () const |
| |
| PNatAAOptERotamerDataOPs::const_iterator | rotamer_data_begin () const |
| |
| PNatAAOptERotamerDataOPs::const_iterator | rotamer_data_end () const |
| |
| | OptEPositionData () |
| |
| virtual | ~OptEPositionData () |
| |
| void | tag (std::string const &tag_in) |
| |
| std::string const & | tag () const |
| |
|
| void | process_rotamers (Multivec const &vars, Size const num_energy_dofs, EnergyMap const &fixed_terms, ScoreTypes const &score_list, ScoreTypes const &fixed_score_list, Size const aa_range, utility::vector1< Real > const &dummy_set, utility::vector1< Real > &best_energy_by_aa, utility::vector1< utility::vector1< Real > > &unweighted_E_dof, Multivec &ref_deriv_weight) const |
| | used by derived class as well – finds the energies for the best rotamer for each amino acid More...
|
| |
| void | update_range (SingleStructureDataCOP structure, ScoreTypes const &free_score_list, ScoreTypes const &fixed_score_list, EnergyMap &lower_bound, EnergyMap &upper_bound) const |
| | Helper function for range(); updates lower/upper_bound as needed so that score_list scores from structure are included in the range. More...
|
| |
Does actual work for OptE minimization
Determine the metric for optimization of energy weights. Original source is Brian Kuhlman's FORTRAN weight training program. Since the goal is to maximize site-by-site sequence recovery, the metric is decomposable by positions, and this does the work for one position. First, a pass is made through all the rotamers at the position, and the lowest energy rotamer for each amino acid is identified. Next, a rough partition function is constructed. The score to be minimized is the negative log of the probabilty of selecting a native aa rotamer (needn't be the native conformation).
Limitations:
- Assumes that the choices available at each position are the 20 canonical amino acids - no DNA base recovery, no pTyr.
- Doesn't have Chris Saunder's compositional constraints. This would require implementation of a direction-set ( aka powell ) minimizer, as his optimization metric doesn't yield gradients.
- Parameters
-
| num_energy_dofs | Basically, turn over all the private data from OptEMultiFunc |
Reimplemented from protocols::optimize_weights::PNatAAOptEPositionData.
References core::chemical::num_canonical_aas, and protocols::optimize_weights::pssm_data.


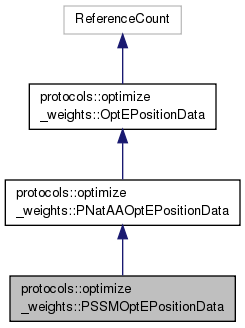
 Public Types inherited from protocols::optimize_weights::PNatAAOptEPositionData
Public Types inherited from protocols::optimize_weights::PNatAAOptEPositionData Public Types inherited from protocols::optimize_weights::OptEPositionData
Public Types inherited from protocols::optimize_weights::OptEPositionData Public Member Functions inherited from protocols::optimize_weights::PNatAAOptEPositionData
Public Member Functions inherited from protocols::optimize_weights::PNatAAOptEPositionData Public Member Functions inherited from protocols::optimize_weights::OptEPositionData
Public Member Functions inherited from protocols::optimize_weights::OptEPositionData Protected Member Functions inherited from protocols::optimize_weights::PNatAAOptEPositionData
Protected Member Functions inherited from protocols::optimize_weights::PNatAAOptEPositionData Protected Member Functions inherited from protocols::optimize_weights::OptEPositionData
Protected Member Functions inherited from protocols::optimize_weights::OptEPositionData 1.8.7
1.8.7