 |
Rosetta
2020.11
|
 |
Rosetta
2020.11
|
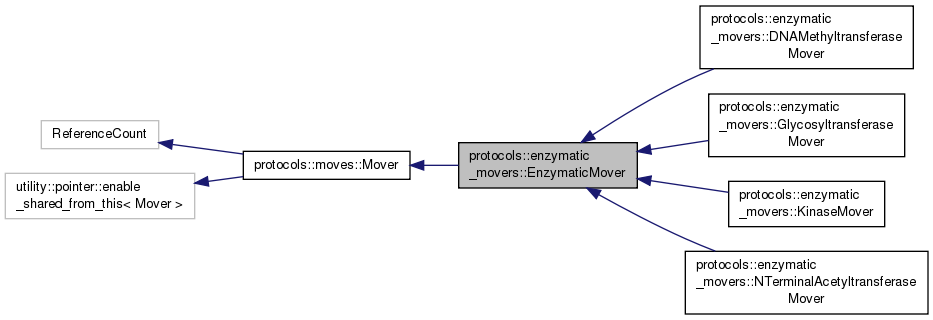
#include <EnzymaticMover.hh>
Public Member Functions | |
| EnzymaticMover () | |
| Default constructor. More... | |
| EnzymaticMover (std::string const &enzyme_family) | |
| Constructor with enzyme family provided. More... | |
| EnzymaticMover (EnzymaticMover const &)=default | |
| Copy constructor. More... | |
| ~EnzymaticMover () override=default | |
| void | show (std::ostream &output=std::cout) const override |
| Generate string representation of EnzymaticMover for debugging purposes. More... | |
| std::string | get_name () const override=0 |
| Return the name of the Mover. More... | |
| moves::MoverOP | clone () const override=0 |
| Return a clone of the Mover object. More... | |
| moves::MoverOP | fresh_instance () const override=0 |
| Generates a new Mover object freshly created with the default ctor. More... | |
| void | parse_my_tag (TagCOP tag, basic::datacache::DataMap &data, Filters_map const &, moves::Movers_map const &, core::pose::Pose const &pose) override |
| Called by MoverFactory when constructing new Movers. Takes care of the specific mover's parsing. More... | |
| void | apply (core::pose::Pose &input_pose) override |
| Apply the corresponding move to <input_pose>. More... | |
| bool | mover_provides_citation_info () const override |
| Does this EnzymaticMover provide information about how to cite it? More... | |
| utility::vector1 < basic::citation_manager::UnpublishedModuleInfoCOP > | provide_authorship_info_for_unpublished () const override |
| Provide a list of authors and their e-mail addresses, as strings. More... | |
| std::string | get_enzyme_family () const |
| Get the family name of this simulated enzyme. More... | |
| void | set_species (std::string const &species_name) |
| Set the species name of this simulated enzyme. More... | |
| std::string | get_species () const |
| Get the species name of this simulated enzyme. More... | |
| void | set_enzyme (std::string const &enzyme_name) |
| Set the specific name of this simulated enzyme. More... | |
| std::string | get_enzyme () const |
| Return the specific name of this simulated enzyme. More... | |
| void | set_efficiency (core::Real setting) |
| Directly set the efficiency of this enzyme, ignoring whatever is in the database. More... | |
| core::Real | get_efficiency () const |
| Get the efficiency of this enzyme. More... | |
| void | exclude_site (core::uint seqpos) |
| Do not perform a reaction at this site, even if it is within a consensus sequence match for this enzyme. More... | |
| void | set_excluded_sites (utility::vector1< core::uint > const &excluded_sites) |
| Pass a list of sequence positions that are forbidden from being modified, even if they are within a consensus sequence match for this enzyme. More... | |
| utility::vector1< core::uint > | get_excluded_sites () const |
| Return a list of sequence positions that are forbidden from being modified, even if they are within a consensus sequence match for this enzyme. More... | |
| void | ensure_site (core::uint seqpos) |
| Definitely modify this site, if it is within a consensus sequence match for this enzyme. More... | |
| void | set_ensured_sites (utility::vector1< core::uint > const &ensured_sites) |
| Pass a list of sequence positions that are guaranteed to be modified, if they are within a consensus sequence match for this enzyme. More... | |
| utility::vector1< core::uint > | get_ensured_sites () const |
| Return a list of sequence positions that are guaranteed to be modified, if they are within a consensus sequence match for this enzyme. More... | |
| core::Size | get_n_reactive_sites () const |
| Return the current number of reactive sites. More... | |
| core::uint | get_reactive_site_sequence_position (core::uint const index) const |
| Return the sequence position of the requested reactive site. More... | |
| std::string const & | get_reactive_site_atom_name (core::uint const index) const |
| Return the atom name of the requested reactive site. More... | |
| core::Size | get_n_co_substrates () const |
| Return the current number of reactive sites. More... | |
| std::string const & | get_co_substrate (core::uint const index) const |
| Return the requested cosubstrate of this enzymatic reaction. More... | |
| void | perform_major_reaction_only () |
| Set this EnzymaticMover to perform only its major reaction. More... | |
| void | perform_all_reactions () |
| Allow this EnzymaticMover to be promiscuous, performing a random transfer from among its possible co-substrates. More... | |
| bool | performs_major_reaction_only () const |
| Does this enzyme only perform its major reaction? More... | |
| void | set_pose_reactive_sites (core::pose::Pose const &pose) |
 Public Member Functions inherited from protocols::moves::Mover Public Member Functions inherited from protocols::moves::Mover | |
| Mover () | |
| virtual MoverOP | create () |
| MoverCOP | get_self_ptr () const |
| MoverOP | get_self_ptr () |
| MoverCAP | get_self_weak_ptr () const |
| MoverAP | get_self_weak_ptr () |
| Mover (std::string const &type_name) | |
| sets the type for a mover; name_ has been removed (2010/05/14) More... | |
| virtual void | test_move (Pose &pose) |
| : Unit test support function. Apply one move to a given pose. Allows extra test specific functions to be called before applying More... | |
| virtual bool | reinitialize_for_each_job () const |
| Inform the Job Distributor (August '08 vintage) whether this object needs to be freshly regenerated on each use. More... | |
| virtual bool | reinitialize_for_new_input () const |
| Inform the Job Distributor (August '08 vintage) whether this object needs to be regenerated when the input pose is about to change, (for example, if the Mover has special code on the first apply() that is only valid for that one input pose). More... | |
| MoverStatus | get_last_move_status () const |
| end parser interface, start Job Distributor interface///////////// More... | |
| void | reset_status () |
| resets status to SUCCESS, meant to be used before an apply(). The job distributor (august 08 vintage) uses this to ensure non-accumulation of status across apply()s. More... | |
| virtual core::pose::PoseOP | get_additional_output () |
| Mechanism by which a mover may return multiple output poses from a single input pose. More... | |
| void | set_type (std::string const &setting) |
| Set the 'type' string. More... | |
| std::string | get_type () const |
| void | type (const std::string &type_in) |
| Set the 'type' string. More... | |
| std::string const & | type () const |
| Get the set 'type' string. More... | |
| virtual void | set_input_pose (PoseCOP pose) |
| setter for poses contained for rms More... | |
| virtual void | set_native_pose (PoseCOP pose) |
| setter for native poses contained for rms -— we should get rid of this method? it is widely used, but a bit unsafe More... | |
| PoseCOP | get_input_pose () const |
| PoseCOP | get_native_pose () const |
| void | set_current_job (protocols::jobdist::BasicJobCOP job) |
| jobdist::BasicJobCOP | get_current_job () const |
| virtual void | set_current_tag (std::string const &new_tag) |
| std::string | get_current_tag () const |
| A tag is a unique identifier used to identify structures produced by this Mover. get_current_tag() returns the tag, and set_current_tag( std::string tag ) sets the tag. This functionality is not intended for use with the 2008 job distributor. More... | |
| virtual core::Real | last_proposal_density_ratio () |
| virtual void | clear_info () |
| Strings container can be used to return miscellaneous info (as std::string) from a mover, such as notes about the results of apply(). The job distributor (Apr 09 vintage) will check this function to see if your protocol wants to add string info to the Job that ran this mover. One way this can be useful is that later, a JobOutputter may include/append this info to an output file. More... | |
| virtual Strings & | info () |
| non-const accessor More... | |
| virtual Strings const & | info () const |
| const accessor More... | |
| virtual utility::vector1 < basic::citation_manager::CitationCollectionCOP > | provide_citation_info () const |
| Provide the citation. More... | |
| virtual bool | mover_is_unpublished () const |
| Does this mover indicate that it is unpublished (and, by extension, that the author should be included in publications resulting from it)? More... | |
Static Public Member Functions | |
| static void | register_options () |
| Register options with the option system. More... | |
| static utility::tag::XMLSchemaComplexTypeGeneratorOP | xml_schema_complex_type_generator () |
 Static Public Member Functions inherited from protocols::moves::Mover Static Public Member Functions inherited from protocols::moves::Mover | |
| static std::string | name () |
| static void | register_options () |
| Overload this static method if you access options within the mover. More... | |
Protected Member Functions | |
| void | set_enzyme_family (std::string const &family_name) |
| Set the family name of this simulated enzyme. More... | |
| virtual void | perform_reaction (core::pose::Pose &input_pose, core::uint const site, std::string const &cosubstrate="")=0 |
| Actually perform the virtual reaction for this specific EnzymaticMover. <input_pose>: The Pose to be acted on. <site>: An index for the reactive site to be acted on. <cosubstrate>: A string providing information to the Enzymatic Mover to add or subtract the proper atoms or Residues from the Pose. How this parameter is handled is specific to each distinct EnzymaticMover. More... | |
 Protected Member Functions inherited from protocols::moves::Mover Protected Member Functions inherited from protocols::moves::Mover | |
| void | set_last_move_status (MoverStatus status) |
| nonvirtual setter for MoverStatus last_status_. Protected means that only the mover itself will be able to change its own status. The job distributor (august 08 vintage) is aware of status set with this function and will do what the MoverStatus says. More... | |
Private Member Functions | |
| void | set_commandline_options () |
| void | init (std::string const &enzyme_family) |
| void | set_efficiency () |
| void | set_available_co_substrates () |
Private Attributes | |
| std::string | enzyme_family_ = "" |
| std::string | species_name_ = "" |
| std::string | enzyme_name_ = "" |
| core::Real | efficiency_ = 1.0 |
| utility::vector1< core::uint > | excluded_sites_ |
| utility::vector1< core::uint > | ensured_sites_ |
| utility::vector1< ReactionSite > | reaction_sites_ |
| utility::vector1< std::string > | co_substrates_ |
| bool | performs_major_reaction_only_ = false |
Additional Inherited Members | |
 Public Types inherited from protocols::moves::Mover Public Types inherited from protocols::moves::Mover | |
| typedef utility::tag::TagCOP | TagCOP |
| typedef core::pose::Pose | Pose |
| typedef core::pose::PoseCOP | PoseCOP |
| typedef protocols::filters::Filters_map | Filters_map |
| typedef std::list< std::string > | Strings |
This is a base class for any Mover that modifies the sequence of a Pose in a way that simulates the action of an enzyme. Any Mover inheriting from this base class must provide an enzyme family corresponding to a directory of enzyme data and must implement the perform_reaction() method, which modifies, adds, or removes (a) Residue(s). The core machinery of this base class uses the enzymatic data to search for potential reaction sites.
| protocols::enzymatic_movers::EnzymaticMover::EnzymaticMover | ( | ) |
Default constructor.
References init().
| protocols::enzymatic_movers::EnzymaticMover::EnzymaticMover | ( | std::string const & | enzyme_family | ) |
Constructor with enzyme family provided.
References init().
|
default |
Copy constructor.
|
overridedefault |
|
overridevirtual |
Apply the corresponding move to <input_pose>.
When applied, every EnzymaticMover first obtains a list of all possible reaction sites, using enzyme data. Next, it loops through the sites and checks if they are ensured. If not, it decides whether or not to modify the site based on the enzymes efficiency, as recorded in the database. Then it performs the specific "reaction" of the particular EnzymaticMover at that site.
| <input_pose> | the structure to be post-translationally modified, i.e., "substrate 1" |
Implements protocols::moves::Mover.
References protocols::sparta::contains(), protocols::moves::FAIL_DO_NOT_RETRY, get_co_substrate(), get_efficiency(), get_ensured_sites(), get_enzyme(), get_n_co_substrates(), get_n_reactive_sites(), get_reactive_site_sequence_position(), perform_reaction(), performs_major_reaction_only(), core::scoring::rg, protocols::moves::Mover::set_last_move_status(), set_pose_reactive_sites(), show(), and protocols::TR().
|
overridepure virtual |
Return a clone of the Mover object.
clone is meant to return an OP'ed deep copy of this object. This really should be a pure virtual in the base class, but adding pure virtuals to Mover would massively disrupt the code. This default implementation crashes at runtime instead of compiletime if you try to call it. If this code is causing you problems, your Mover needs to override this function.
Reimplemented from protocols::moves::Mover.
Implemented in protocols::enzymatic_movers::DNAMethyltransferaseMover, protocols::enzymatic_movers::GlycosyltransferaseMover, protocols::enzymatic_movers::KinaseMover, and protocols::enzymatic_movers::NTerminalAcetyltransferaseMover.
| void protocols::enzymatic_movers::EnzymaticMover::ensure_site | ( | core::uint | seqpos | ) |
Definitely modify this site, if it is within a consensus sequence match for this enzyme.
References ensured_sites_.
| void protocols::enzymatic_movers::EnzymaticMover::exclude_site | ( | core::uint | seqpos | ) |
Do not perform a reaction at this site, even if it is within a consensus sequence match for this enzyme.
References excluded_sites_.
|
overridepure virtual |
Generates a new Mover object freshly created with the default ctor.
fresh_instance is meant to return a new object of this class, created with the default constructor. This really should be a pure virtual in the base class, but adding pure virtuals to Mover would massively disrupt the code. This default implementation crashes at runtime instead of compiletime if you try to call it. If this code is causing you problems, your Mover needs to override this function. This is used by the August 08 job distributor.
Reimplemented from protocols::moves::Mover.
Implemented in protocols::enzymatic_movers::DNAMethyltransferaseMover, protocols::enzymatic_movers::GlycosyltransferaseMover, protocols::enzymatic_movers::KinaseMover, and protocols::enzymatic_movers::NTerminalAcetyltransferaseMover.
|
inline |
Return the requested cosubstrate of this enzymatic reaction.
References co_substrates_.
Referenced by apply().
|
inline |
Get the efficiency of this enzyme.
An efficiency of 0.45 means that this enzyme will perform its reaction at any recognized site 45% of the time.
References efficiency_.
Referenced by apply().
|
inline |
Return a list of sequence positions that are guaranteed to be modified, if they are within a consensus sequence match for this enzyme.
References ensured_sites_.
Referenced by apply().
|
inline |
|
inline |
Get the family name of this simulated enzyme.
This EnzymaticMover is a member of this enyzme family.
References enzyme_family_.
|
inline |
Return a list of sequence positions that are forbidden from being modified, even if they are within a consensus sequence match for this enzyme.
References excluded_sites_.
|
inline |
Return the current number of reactive sites.
References co_substrates_.
|
inline |
|
overridepure virtual |
Return the name of the Mover.
Implements protocols::moves::Mover.
Implemented in protocols::enzymatic_movers::DNAMethyltransferaseMover, protocols::enzymatic_movers::GlycosyltransferaseMover, protocols::enzymatic_movers::KinaseMover, and protocols::enzymatic_movers::NTerminalAcetyltransferaseMover.
Referenced by provide_authorship_info_for_unpublished().
|
inline |
Return the atom name of the requested reactive site.
References reaction_sites_.
Referenced by protocols::enzymatic_movers::GlycosyltransferaseMover::perform_reaction().
|
inline |
Return the sequence position of the requested reactive site.
References reaction_sites_.
Referenced by apply(), protocols::enzymatic_movers::NTerminalAcetyltransferaseMover::perform_reaction(), protocols::enzymatic_movers::DNAMethyltransferaseMover::perform_reaction(), protocols::enzymatic_movers::GlycosyltransferaseMover::perform_reaction(), and protocols::enzymatic_movers::KinaseMover::perform_reaction().
|
inline |
Get the species name of this simulated enzyme.
This EnzymaticMover is limited to reactions known to occur in the returned species.
References species_name_.
|
private |
References set_commandline_options(), set_enzyme_family(), and protocols::TR().
Referenced by EnzymaticMover().
|
overridevirtual |
Does this EnzymaticMover provide information about how to cite it?
Reimplemented from protocols::moves::Mover.
Reimplemented in protocols::enzymatic_movers::GlycosyltransferaseMover, and protocols::enzymatic_movers::KinaseMover.
|
overridevirtual |
Called by MoverFactory when constructing new Movers. Takes care of the specific mover's parsing.
Some movers need not be parsed, so we shouldn't stop executions. This, however, calls attention to the lack of this method, which could be due to something as silly as a wrong parameters definition.
Reimplemented from protocols::moves::Mover.
References perform_all_reactions(), perform_major_reaction_only(), set_efficiency(), set_enzyme(), and set_species().
|
inline |
Allow this EnzymaticMover to be promiscuous, performing a random transfer from among its possible co-substrates.
References performs_major_reaction_only_.
Referenced by parse_my_tag().
|
inline |
Set this EnzymaticMover to perform only its major reaction.
References performs_major_reaction_only_.
Referenced by parse_my_tag().
|
protectedpure virtual |
Actually perform the virtual reaction for this specific EnzymaticMover. <input_pose>: The Pose to be acted on. <site>: An index for the reactive site to be acted on. <cosubstrate>: A string providing information to the Enzymatic Mover to add or subtract the proper atoms or Residues from the Pose. How this parameter is handled is specific to each distinct EnzymaticMover.
Implemented in protocols::enzymatic_movers::GlycosyltransferaseMover, protocols::enzymatic_movers::KinaseMover, protocols::enzymatic_movers::DNAMethyltransferaseMover, and protocols::enzymatic_movers::NTerminalAcetyltransferaseMover.
Referenced by apply().
|
inline |
Does this enzyme only perform its major reaction?
References performs_major_reaction_only_.
Referenced by apply().
|
overridevirtual |
Provide a list of authors and their e-mail addresses, as strings.
Reimplemented from protocols::moves::Mover.
Reimplemented in protocols::enzymatic_movers::GlycosyltransferaseMover, and protocols::enzymatic_movers::KinaseMover.
References get_name().
|
static |
Register options with the option system.
References protocols::moves::Mover::register_options().
Referenced by protocols::enzymatic_movers::DNAMethyltransferaseMover::register_options(), protocols::enzymatic_movers::NTerminalAcetyltransferaseMover::register_options(), protocols::enzymatic_movers::KinaseMover::register_options(), and protocols::enzymatic_movers::GlycosyltransferaseMover::register_options().
|
private |
References co_substrates_, enzyme_family_, enzyme_name_, core::enzymes::EnzymeManager::get_second_substrates_or_byproducts(), and species_name_.
Referenced by set_enzyme(), and set_species().
|
private |
References set_efficiency(), set_enzyme(), and set_species().
Referenced by init().
|
inline |
Directly set the efficiency of this enzyme, ignoring whatever is in the database.
An efficiency of 0.45 means that this enzyme will perform its reaction at any recognized site 45% of the time.
References efficiency_.
|
private |
References efficiency_, enzyme_family_, enzyme_name_, core::enzymes::EnzymeManager::get_efficiency(), and species_name_.
Referenced by parse_my_tag(), set_commandline_options(), set_enzyme(), and set_species().
|
inline |
Pass a list of sequence positions that are guaranteed to be modified, if they are within a consensus sequence match for this enzyme.
References ensured_sites_.
| void protocols::enzymatic_movers::EnzymaticMover::set_enzyme | ( | std::string const & | enzyme_name | ) |
Set the specific name of this simulated enzyme.
If set, this EnzymaticMover will use specific enzymatic details for this reaction from the database. If the species name has not been set, an enzyme from "h_sapiens" is assumed.
| <setting> | An enzyme name as listed in an appropriate enzyme file in "database/virtual_enzymes/glycosyltransferases/<species_name_>/" |
References enzyme_name_, set_available_co_substrates(), set_efficiency(), and species_name_.
Referenced by parse_my_tag(), and set_commandline_options().
|
protected |
Set the family name of this simulated enzyme.
This method is protected; it should only ever be called by a derived class.
References enzyme_family_.
Referenced by init().
|
inline |
Pass a list of sequence positions that are forbidden from being modified, even if they are within a consensus sequence match for this enzyme.
References excluded_sites_.
| void protocols::enzymatic_movers::EnzymaticMover::set_pose_reactive_sites | ( | core::pose::Pose const & | pose | ) |
| void protocols::enzymatic_movers::EnzymaticMover::set_species | ( | std::string const & | species_name | ) |
Set the species name of this simulated enzyme.
Setting the species name limits the behavior of this EnzymaticMover to reactions known to occur in the given species.
| <setting> | A species name in the format "e_coli" or "h_sapiens", which must correspond to a directory in the Rosetta database, e.g., "database/virtual_enzymes/glycosyltransferases/h_sapiens/" |
References enzyme_name_, set_available_co_substrates(), set_efficiency(), and species_name_.
Referenced by parse_my_tag(), and set_commandline_options().
|
overridevirtual |
Generate string representation of EnzymaticMover for debugging purposes.
Reimplemented from protocols::moves::Mover.
References co_substrates_, efficiency_, ensured_sites_, enzyme_name_, excluded_sites_, get_n_co_substrates(), performs_major_reaction_only_, protocols::moves::Mover::show(), and species_name_.
Referenced by apply(), and protocols::enzymatic_movers::operator<<().
|
static |
References protocols::moves::complex_type_name_for_mover().
Referenced by protocols::enzymatic_movers::DNAMethyltransferaseMover::provide_xml_schema(), protocols::enzymatic_movers::NTerminalAcetyltransferaseMover::provide_xml_schema(), protocols::enzymatic_movers::KinaseMover::provide_xml_schema(), and protocols::enzymatic_movers::GlycosyltransferaseMover::provide_xml_schema().
|
private |
Referenced by get_co_substrate(), get_n_co_substrates(), set_available_co_substrates(), and show().
|
private |
Referenced by get_efficiency(), set_efficiency(), and show().
|
private |
Referenced by ensure_site(), get_ensured_sites(), set_ensured_sites(), and show().
|
private |
Referenced by get_enzyme_family(), set_available_co_substrates(), set_efficiency(), set_enzyme_family(), and set_pose_reactive_sites().
|
private |
Referenced by get_enzyme(), set_available_co_substrates(), set_efficiency(), set_enzyme(), set_pose_reactive_sites(), set_species(), and show().
|
private |
Referenced by exclude_site(), get_excluded_sites(), set_excluded_sites(), set_pose_reactive_sites(), and show().
|
private |
Referenced by perform_all_reactions(), perform_major_reaction_only(), performs_major_reaction_only(), and show().
|
private |
|
private |
Referenced by get_species(), set_available_co_substrates(), set_efficiency(), set_enzyme(), set_pose_reactive_sites(), set_species(), and show().
 1.8.7
1.8.7