Ultra-large virtual screening using a motif-guided Rosetta pipeline validated in zebrafish
Broader significance: a screenable path from computation to in vivo
Virtual screening can now evaluate billions of molecules. But that scale creates a practical problem. Docking pipelines often return thousands of candidates that pass computational filters, and predicted binding energies do not reliably predict what will work in living systems. This study introduces a workflow designed to better prioritize hits computationally and then test them in a whole animal.
Background: the bottleneck after docking
Most docking approaches rely heavily on energy scoring. Because these scoring functions balance speed and accuracy, predicted binding energies correlate only loosely with experimental affinity. As a result, researchers may manually inspect thousands of molecules before deciding which to synthesize. Even strong binders can fail in animals due to off-target effects or biological barriers.
The authors aimed to reduce subjective triage and pair improved computational prioritization with scalable in vivo testing.
The new approach: REAL-M, guided by structural interaction motifs
REAL-M, the Rosetta Engine for Anchoring Ligands with a Motif, is built within the Rosetta modeling suite. Instead of relying solely on docking scores, it uses structural interaction patterns extracted from almost 222,000 protein–ligand complexes in the Protein Data Bank. These “motifs” encode recurring geometric relationships between amino acids and ligand atoms and are used to guide ligand placement relative to a selected anchor residue.
To scale to ultra-large libraries, the authors expanded 2.6 billion Enamine REAL molecules to 34.6 billion conformers. Shape filtering reduced this space before motif-guided docking. Additional filters, including a rapid grid-based clash check and post-docking motif screening, limited the number of candidates entering high-resolution refinement.
Key results: benchmarking and hypocretin receptor hits
REAL-M was benchmarked on 84 protein–ligand systems. Its low-energy placements were reported as consistently comparable to or more accurate than several widely used docking programs. In a separate recovery test, REAL-M rediscovered 34 of 36 ligands from prior docking campaigns when each was embedded among 10,000 similarly sized models.
For prospective discovery, the authors targeted hypocretin receptor 2, a GPCR involved in sleep regulation and conserved between humans and zebrafish. After filtering and manual review, 95 molecules were selected for synthesis, and 82 were tested in a GPCR assay.
More than half significantly inhibited binding of the orexin A peptide. Of compounds docked to the inactive receptor structure, 28 of 30 were significant antagonists. Of those docked to the active structure, 21 of 52 were also effective antagonists. The seven strongest inhibitors showed potency comparable to commercially available antagonists.
Validation: structural controls and zebrafish testing
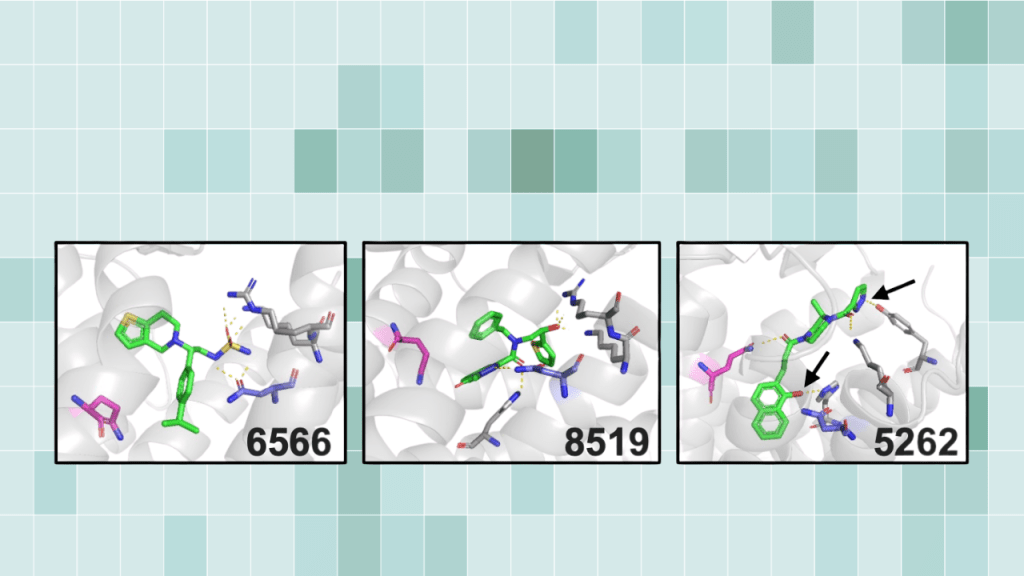
To support one predicted binding mode, the authors synthesized two analogs missing atoms involved in a modeled interaction. Both analogs eliminated antagonist activity.
The top seven antagonists were then tested in zebrafish. In a transgenic line where hypocretin overexpression induces hyperactivity, three compounds significantly mitigated the behavioral phenotype. A zebrafish hcrtr2 loss-of-function mutant was generated to assess specificity. Known antagonists such as suvorexant reduced movement even in mutants, suggesting off-target effects. In contrast, most newly identified compounds showed only mild, non-significant effects in mutants.
Broader significance: a screenable path from computation to in vivo
The authors position zebrafish as a practical intermediate validation system between cell assays and mammalian models. Behavioral experiments required relatively small amounts of compound and enabled direct testing of functional effects in a whole organism.
They also report that among 210 GPCRs examined, 165 have binding pockets more than 90 percent similar between human and zebrafish. This conservation supports the feasibility of applying the combined motif-guided docking and zebrafish validation strategy to other conserved receptor targets.
This work was recently posted as a bioRxiv preprint by Thyme, Ginsparg, and colleagues.
