Blog post
Registrations are now open for European RosettaCon 2026
The registrations for the 5th European RosettaCon – Crossing Boundaries with Protein Design, taking place October 28–30, 2026 at the NOVA University Lisbon (Portugal) are now open! The meeting will feature invited talks, contributed presentations, and poster sessions, offering a unique opportunity to share discoveries, connect with colleagues, and foster collaborations within the international protein…
Read MoreSummer RosettaCon 2026 registration is now open
We are pleased to announce that Summer RosettaCon 2026 will take place at the University of Washington in Seattle from August 3–7, 2026. The conference will begin Monday afternoon at 5:00 p.m. and conclude Thursday evening. Friday will be the traditional hike/activity day, beginning after breakfast. Registration Registration is NOW OPEN. Early registration pricing is…
Read MoreCyclicMPNN adapts sequence design to cyclic backbones
Why this work matters Cyclic peptides are attractive scaffolds because their closed backbone can promote structural rigidity and proteolytic stability. Yet assigning sequences that reliably fold into a given cyclic backbone remains a key challenge. In this study, the authors introduce CyclicMPNN, a fine-tuned version of ProteinMPNN designed specifically to improve sequence design for short…
Read MoreAngela Colbert joins Rosetta Commons as Communications Lead
Angela Colbert, Ph.D., joined us on February 23 as our new Communications Lead. She comes to us from NASA’s Jet Propulsion Laboratory, where she spent the past five years leading science communication initiatives for major Earth science efforts. In that role, she translated complex climate and satellite data into clear, high-impact stories across web, multimedia,…
Read MoreFoundry Docker Image Now Available
Rosetta Commons now offers an official Foundry Docker image designed to make it dramatically easier to run cutting-edge biomolecular machine learning models without complex setup. Foundry is the central repository for a suite of ML models used in protein science—including RF3 (a next-generation biomolecular structure prediction network), ProteinMPNN inverse-folding models, and the newest RFdiffusion3 (RFD3)…
Read MoreEngineering a protein biosensor to detect common NSAIDs in wastewater
Why this research matters Pharmaceuticals often end up in wastewater after use, and monitoring them can be technically demanding. Traditional analytical methods are powerful but typically require specialized equipment and off-site processing, which can limit how frequently water is tested. This study describes a protein-based biosensor designed to detect two widely used “profen” non-steroidal anti-inflammatory…
Read MoreRegistration open: Immunogen design workshop at Vanderbilt
The Meiler Lab at Vanderbilt University will host an in-person workshop, Computationally Guided Immunogen Design with Rosetta and ML Tools, on April 13–15, 2025. The program is intended for academic and industry researchers seeking structured, hands-on training in computational immunogen design. What the workshop covers Over three days, participants will work through guided tutorials focused…
Read MoreDocumentation Playbook Now Available
The Documentation Playbook is a new guide designed to make writing and maintaining documentation for your software projects clearer and easier. The Documentation Playbook is part of the Playbooks Project from the Open Molecular Software Foundation (OMSF). What’s inside: An overview of the different types of documentation that are useful for open-source scientific software projects…
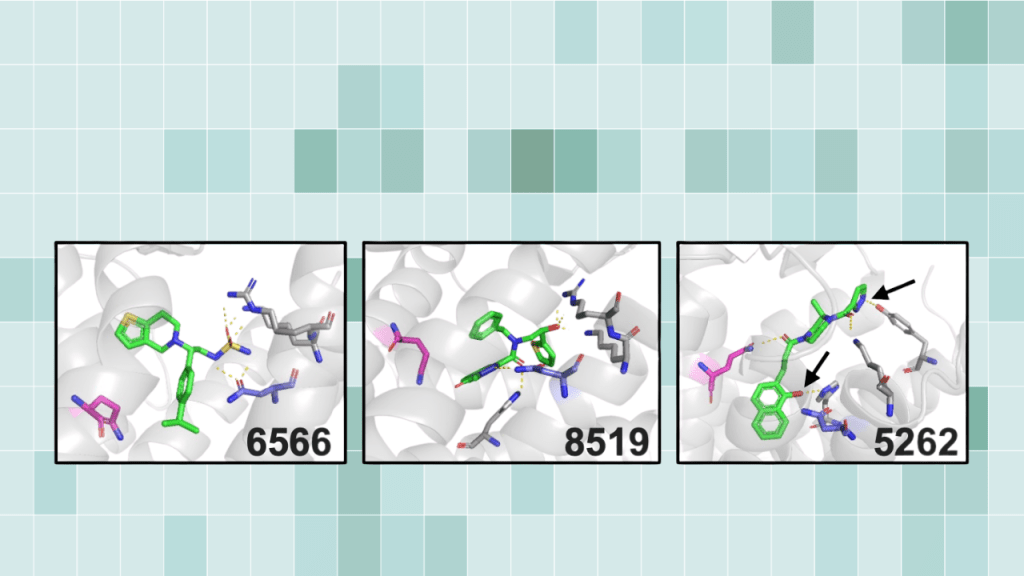
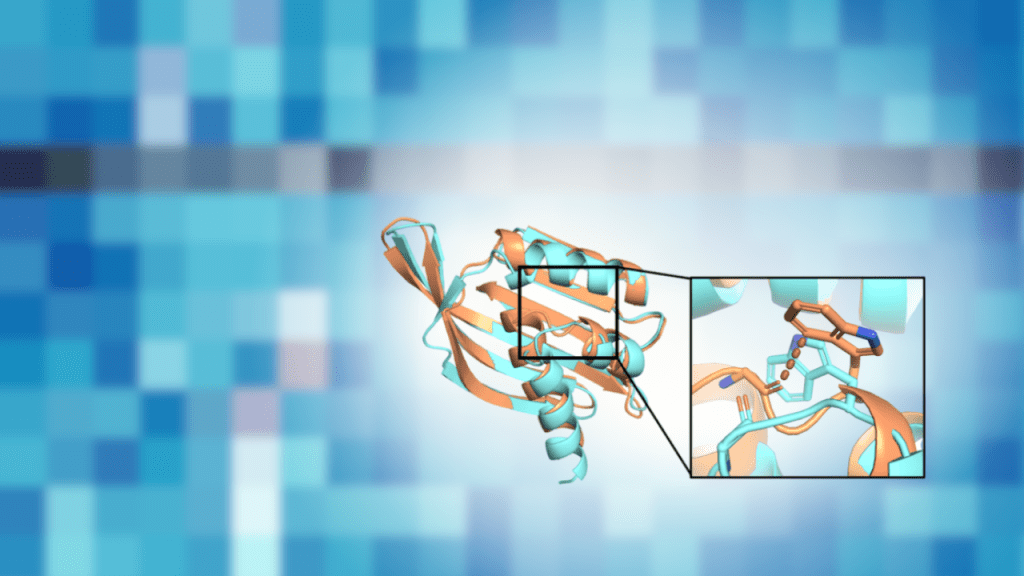
Read MoreUltra-large virtual screening using a motif-guided Rosetta pipeline validated in zebrafish
Broader significance: a screenable path from computation to in vivo Virtual screening can now evaluate billions of molecules. But that scale creates a practical problem. Docking pipelines often return thousands of candidates that pass computational filters, and predicted binding energies do not reliably predict what will work in living systems. This study introduces a workflow…
Read MoreUsing protein design experiments to guide energy-function development
Why this matters When a protein is designed on a computer, tiny “impossible overlaps” between atoms can slip into the model. Those clashes may look small on-screen, but they can push a real protein to adopt a different shape than intended. In a December 2025 bioRxiv preprint, the authors show that steric clashing is a…
Read More