Science Updates
Speed Up PyRosetta Development with Autocompletion and Type Checking
By Harrison Truscott PyRosetta, the Python interface to the Rosetta binary, now comes packaged with type stub files (.pyi). These files describe the module’s classes, methods, and variables, as well as function signatures, types of function parameters and return values, and descriptive docstrings. Type stub files can be used by the Integrated Development Environment (IDE)…
Read MoreA Zero-Shot Approach to de novo Metalloenzyme Design
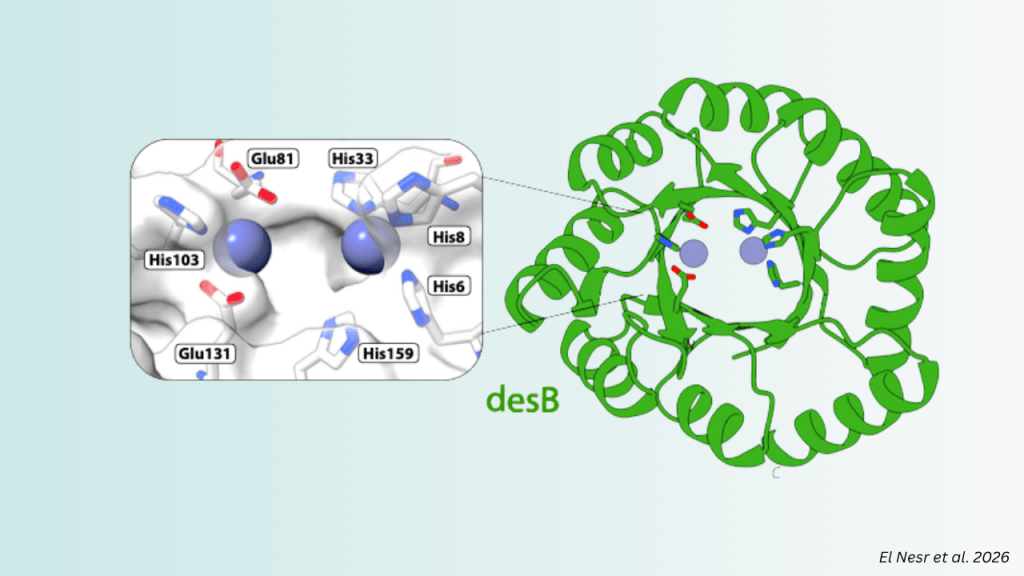
Key Takeaways: dEVA (design by EVolutionary Algorithm) is a framework that achieves the zero-shot design of a highly efficient metalloenzyme without any reliance on natural templates. Design objectives were tailored to the specific chemistry of metalloenzymes, in particular catalytic zinc sites, and from only three experimentally-tested designs found desB, the most efficient de novo designed…
Read MoreDeep Learning’s Impact on de novo Protein Design
Key Takeaways: With advancements in AI, the field of protein engineering has shifted from template-based approaches to de novo designs with approaches including seed-based design, deep generative models, and binder hallucination. Design towards specific protein functions is the current frontier factoring in parameters including neosurfaces, conditional binding with biological stimuli, and dynamic modeling, linking deep…
Read MoreNext Generation Generative Model Unlocks de novo Designs at Scale
Key Takeaways: NVIDIA’s new Proteina-Complexa model combines generative AI with inference-time search algorithms to operate 30-60x faster than RFdiffusion when designing custom proteins that target specific diseases. In a test to find binders for 127 different targets from over 1 million protein designs, the model successfully found binders for 68% of the targets. The model…
Read MoreRosetta Commons Run (rc-run) Now Available
We are pleased to announce a new Rosetta Commons workflow optimization tool: rc-run. rc-run (invoked as rc) is a command-line utility tool for running containerized and native biomolecular software. It simplifies common tasks, such as mounting local directories, building HPC containers, running containerized applications, and logging executed commands, to make complex workflows easier to reliably…
Read MoreCyclicMPNN adapts sequence design to cyclic backbones
Why this work matters Cyclic peptides are attractive scaffolds because their closed backbone can promote structural rigidity and proteolytic stability. Yet assigning sequences that reliably fold into a given cyclic backbone remains a key challenge. In this study, the authors introduce CyclicMPNN, a fine-tuned version of ProteinMPNN designed specifically to improve sequence design for short…
Read MoreFoundry Docker Image Now Available
Rosetta Commons now offers an official Foundry Docker image designed to make it dramatically easier to run cutting-edge biomolecular machine learning models without complex setup. Foundry is the central repository for a suite of ML models used in protein science—including RF3 (a next-generation biomolecular structure prediction network), ProteinMPNN inverse-folding models, and the newest RFdiffusion3 (RFD3)…
Read MoreEngineering a protein biosensor to detect common NSAIDs in wastewater
Why this research matters Pharmaceuticals often end up in wastewater after use, and monitoring them can be technically demanding. Traditional analytical methods are powerful but typically require specialized equipment and off-site processing, which can limit how frequently water is tested. This study describes a protein-based biosensor designed to detect two widely used “profen” non-steroidal anti-inflammatory…
Read MoreUltra-large virtual screening using a motif-guided Rosetta pipeline validated in zebrafish
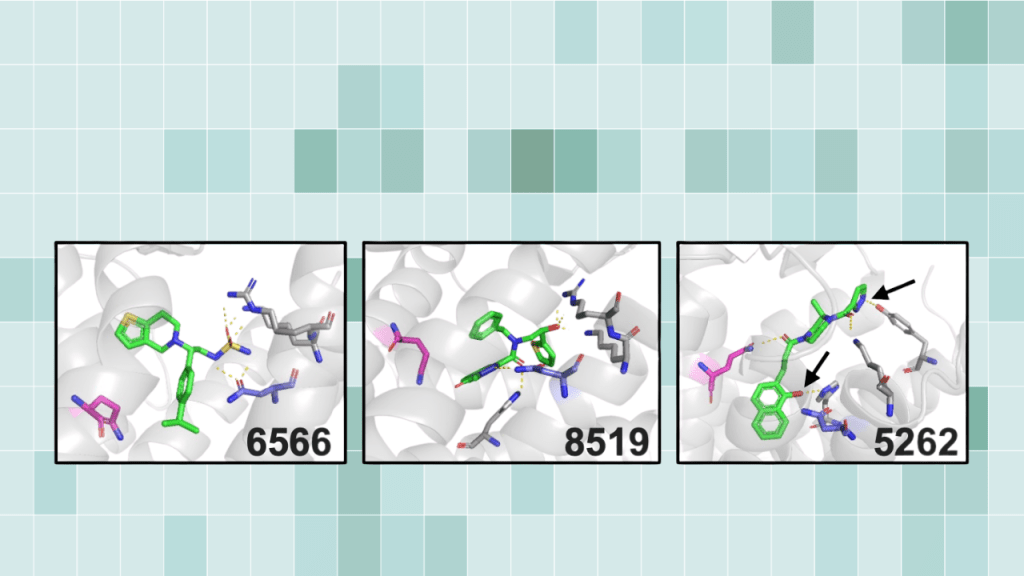
Broader significance: a screenable path from computation to in vivo Virtual screening can now evaluate billions of molecules. But that scale creates a practical problem. Docking pipelines often return thousands of candidates that pass computational filters, and predicted binding energies do not reliably predict what will work in living systems. This study introduces a workflow…
Read MoreUsing protein design experiments to guide energy-function development
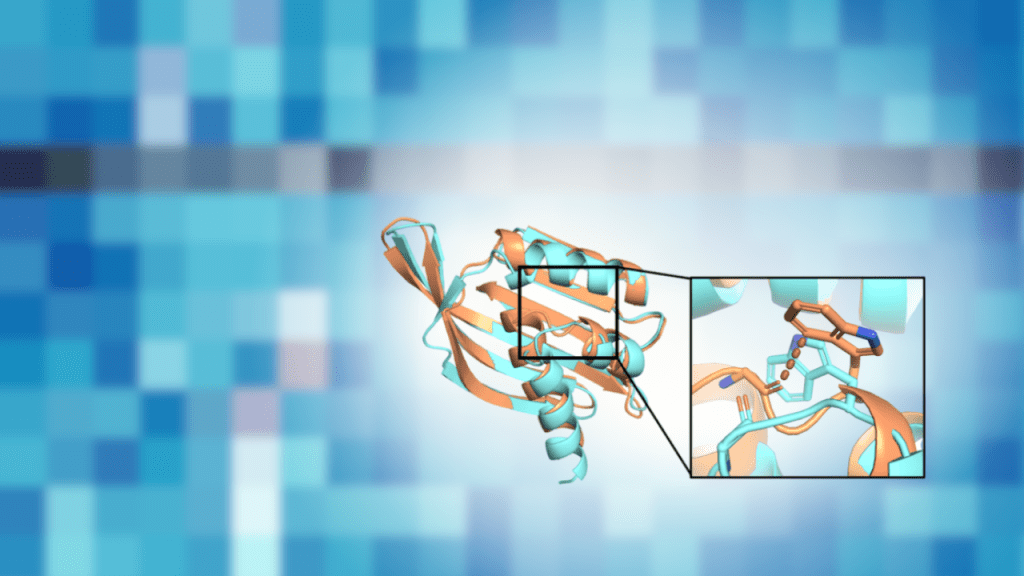
Why this matters When a protein is designed on a computer, tiny “impossible overlaps” between atoms can slip into the model. Those clashes may look small on-screen, but they can push a real protein to adopt a different shape than intended. In a December 2025 bioRxiv preprint, the authors show that steric clashing is a…
Read More